R Short Notes | Cheat-Sheet for Statistical Computing

This is an comprehensive guide to R programming language and its application in the world of Statistics.
This is an intermediate introduction to R and how we can use the programming language to perform data manipulation, analysis and visualisation.
I have learned these stuff from proffesors Arnab Chakraborty and Debasis Sengupta at ISI Kolkata and a lot credit for this goes to them.
I will advise you to run the programs along side reading this to get a better learning experience. Also search the internet for some source files mentioned in the examples, this way you will actualy get your hands dirty and learn things even faster.
Also since it is a pretty large collection of “short-notes”, it is advised that you try to learn through it in a bite-sized format maybe taking several days or weeks to complete.
R (cheatsheet)
Basics
Simple mathematics
R may be used like a simple calculator.
For example:-
3+5 returns 8
Variables
| |
Assignments
=and<-are both used for assignment in the current environment or scope.<<-is used for assignment in parent environments (outside of the current scope), which is useful for global modifications but should be used carefully to avoid unintended side effects.
Standard functions
| |
Vectors
R can handle vectors, which are basically lists of numbers.
Initialising a vector
| |
Entrywise operation on vectors
Most operations that work with numbers act entrywise when applied to vectors.
It is very easy to add/subtract/multiply/divide two vectors entry by entry. Try it yourself.
Summarizing a vector
| |
summary() function is used to obtain a summary of the statistical properties of one or more R objects, output of summary() depends on the class of the object being summarized.
| |
Output:
| Min | 1st Qu. | Median | Mean | 3rd Qu. | Max |
|---|---|---|---|---|---|
| -4.000 | -1.500 | 2.000 | 8.143 | 3.000 | 56.000 |
| |
Extracting parts of a vector
- If x is a vector of length 3 then its entries may be accessed as
x[1],x[2]andx[3]. - Note that the counting starts from 1 and proceeds left-to-right. The quantity inside the square brackets is called the subscript or index.
| |
You can also give negative subscripts.
| |
Comparing vectors
Boolean operators includes: < ,<=, > , >=, ==, !=. You can combine these with, & (and), | (or) and ! (not).(modulo operator %% (e.g., 5%%3 is 2))
Boolean operator %in% which is the “belongs to” operator. For instance, 4 %in% c(3,6,4,1)
returns TRUE.
| |
In R, the all function is used to determine whether all the elements of a given logical vector are TRUE. It returns a single TRUE if all elements are TRUE, and FALSE otherwise.
| |
The any() function in R is used to determine if any elements in a logical vector are TRUE. It returns a single logical value: TRUE if at least one element is TRUE, and FALSE if all elements are FALSE
| |
SORTING
| |
Sometimes we need to order one vector according to another vector.
| |
Notice that ord[1] is the position of the smallest number, ord[2] is the position of the next smallest number, and so on.
| |
Working directory and Libraries
If you are not sure check the current working directory using the getwd() function. You may change the working directory using setwd('path to your desired directory'). Use dir() to List the Files in a Directory/Folder.
To install a library or a package:
install.packages('zyp') # Quotes (single/double) are must!
To load a library in system:
library(zyp) # You may or may not use quotes.
Matrix
| |
| data | an optional data vector (including a list) |
|---|---|
| nrow | the desired number of rows. |
| ncol | the desired number of columns. |
| byrow | logical. If FALSE (the default) the matrix is filled by columns, otherwise the matrix is filled by rows. |
| dimnames | An attribute for the matrix: NULL or a list of length 2 giving the row and column names respectively. An empty list is treated as NULL, and a list of length one as row names. The list can be named, and the list names will be used as names for the dimensions. |
| x | an R object. |
Matrix operations
| |
Now suppose that we want to find the
sum of each column. We use apply function for that.
apply(A,2,sum) # apply the sum function columnwise. The 2 above means columnwise. If we need to find the rowwise means we can use
apply(A,1,mean)
vector vs matrix
Subscripting a matrix is done much like subscripting a vector, except that for a matrix we need two subscripts. To see the (1,2)-th entry (i.e., the entry in row 1 and column 2) of A type; A[1,2]
| |
The as.matrix function converts a vector to a column matrix OR a dataframe to a matrix.
| |
Other matrix functions
diag function
The diag function has two purposes:
When applied to matrices it extracts the diagonal entries.
diag(A)- extracts the diagonal elements of matrixAWhen applied to vectors, it creates a diagonal matrix.
diag(x)- creates a diagonal matrix using vectorx. Sincexhas three elements (1, 2, 3), the resulting diagonal matrix will be a 3x3 matrix with the values ofxon the diagonal and zeros elsewhere.
NOTE:-
diag(x1) # 1
The reason for this behavior is that diag() extracts the diagonal elements from a matrix or creates a diagonal matrix when given a vector. In your case, since x1 is a 1x3 matrix, it effectively returns the single element present at the intersection of the first row and the first column, which is 1
In R, when you apply diag() to a single-row matrix (like your x1), it assumes that you want to extract the elements along the main diagonal of the matrix. In your case, since x1 is a 1x3 matrix (a single row with three columns), it considers the first element (element at the intersection of the first row and the first column) as the main diagonal element.
The behavior is different when you apply diag() to a vector, where it constructs a diagonal matrix with the vector’s elements on the main diagonal.
bind functions
In R, cbind and rbind are functions used for combining data frames or matrices by column or row, respectively. Here’s how they work:
cbind (Column Bind):
The cbind function is used to combine two or more data frames or matrices by adding their columns together. It appends the columns of one data structure to the right of the other
rbind (Row Bind):
The rbind function is used to combine two or more data frames or matrices by adding their rows together. It appends the rows of one data structure below the other
| |
Lists
Unlike a vector or a matrix a list can hold different kinds of objects. Lists are useful in R because they allow R functions to return multiple values.
Example 1:
| |
Example 2:
| |
In R, double square brackets [[]] are used for extracting elements from a list or a data frame. This indexing method allows you to access specific elements or columns within these data structures. If you want to extract elements using integer indices, you can use single square brackets [] with the integer index instead. For example, my_list[[1]] would extract the first element of a list or data frame.
| |
| |
Data Frame
data.frame(..., row.names = NULL, check.rows = FALSE, check.names = TRUE, fix.empty.names = TRUE, stringsAsFactors = FALSE)
The function data.frame() creates data frames, tightly coupled collections of variables which share many of the properties of matrices and of lists, used as the fundamental data structure by most of R’s modeling software.
| parameters | details |
|---|---|
| row.names | NULL or a single integer or character string specifying a column to be used as row names, or a character or integer vector giving the row names for the data frame. |
| check.rows | if TRUE then the rows are checked for consistency of length and names. |
| check.names | logical. If TRUE then the names of the variables in the data frame are checked to ensure that they are syntactically valid variable names and are not duplicated. If necessary they are adjusted (by make.names) so that they are. |
| fix.empty.names | logical indicating if arguments which are “unnamed” (in the sense of not being formally called as someName = arg) get an automatically constructed name or rather name “”. Needs to be set to FALSE even when check.names is false if "" names should be kept. |
| stringsAsFactors | logical: should character vectors be converted to factors? The ‘factory-fresh’ default has been TRUE previously but has been changed to FALSE for R 4.0.0. |
example : -
| |
Arrays
- An array in R can have one, two or more dimensions. It is simply a vector which is stored with additional attributes giving the dimensions (attribute
"dim") and optionally names for those dimensions (attribute"dimnames"). - A two-dimensional array is the same thing as a
matrix. - One-dimensional arrays often look like vectors
- The
"dim"attribute is an integer vector of length one or more containing non-negative values: the product of the values must match the length of the array. - The
"dimnames"attribute is optional: if present it is a list with one component for each dimension, eitherNULLor a character vector of the length given by the element of the"dim"attribute for that dimension. as.arrayis a generic function for coercing to arrays. The default method does so by attaching adimattribute to it. It also attachesdimnamesifxhasnames. The sole purpose of this is to make it possible to access thedim[names]attribute at a later time.is.arrayreturnsTRUEorFALSEdepending on whether its argument is an array (i.e., has adimattribute of positive length) or not.
array(data = NA, dim = length(data), dimnames = NULL) as.array(x, ...) is.array(x)
| parameters | details |
|---|---|
| data | a vector expression vector giving data to fill the array. Non-atomic classed objects are coerced by as.vector. |
| dim | the dim attribute for the array to be created, that is an integer vector of length one or more giving the maximal indices in each dimension. |
| dimnames | either NULL or the names for the dimensions. This must be a list (or it will be ignored) with one component for each dimension, either NULL or a character vector of the length given by dim for that dimension. The list can be named, and the list names will be used as names for the dimensions. If the list is shorter than the number of dimensions, it is extended by NULLs to the length required. |
| x | an R object. |
| |
Control structures
In R, loops allow you to repeat a block of code for each element in a vector or sequence. The for loop is commonly used for this purpose.
The basic syntax of a for loop in R is:
| |
This loop will iterate over each element in someVector, assigning the value of each element to x one by one, and then executing the code inside the loop for each value of x.
EXAMPLE:-
| |
| |
Example use
The following technique can be used whenver you want to create a vector by iterating over something to get something from it.
| |
FUNCTIONS
Definition
Let we have a function, . we can write function to do this as follows:
| |
| |
In body of the function consists of one or more statements, each of which serve one or more of the these three purposes:
internal computation, e.g., the line computes , which is not visible from outside the function.
z = x+yzside-effect, e.g., the line prints a line on the screen.
cat("The difference = ',x-y,'\n')output, i.e., the last line outputs . The last line is always an output line in R. If you want to produce output at any other line, use .
sin(z)return(...)
The input names are dummy variables (has nothing to do with other variables with the same name). More on this later. The inputs are called the arguments or the parameters of the function. The variables outside a function, and those inside a function are linked only via the arguments and returned value. If you are in a desperate need of a function that modifies a variable value outside, then use the <<- assignment instead of =. But use the <<- assignment very sparingly, as its careless use generally leads to bugs that are hard to detect.
Peeping inside the body of a function
A simple way to peep inside the definition of a function is to type the name of the function (sans any parentheses or arguments) at the command prompt, and hit enter. The entire body of the function will be listed on screen.
Functions returning multiple values
| |
Named arguments
The name of an argument may be abbreviated as long as there is no ambiguity.
| |
The cat function is a output function that can print a sequence objects in an unformatted way (unlike print that prints a single object in a formatted way).
print()is designed to print R objects (such as vectors, data frames, and results of expressions) in a user-friendly format. It often adds line breaks and formatting to make the output more readable.print()separates the printed objects with a space by default.cat()is a lower-level function that simply concatenates and prints character strings or other objects without any additional formatting. It doesn’t automatically add line breaks or formatting.cat()allows you to specify a custom separator using thesepargument. By default, it separates the objects with a single space.
Defaults
| |
A default value can be an expression also as shown next.
| |
Optional arguments
| |
The use of ... allows you to pass any valid graphical parameters that the plot() function accepts, giving you flexibility in customizing the appearance of the plot according to your specific requirements.
R as a functional programming language
x = c(3,4,6,1)
x = function(t) t^2
The first makes x an object, while the second makes it a function. R treats objects and functions on an equal footing. You can do with functions pretty much whatever you can do with objects, e.g., you can pass a function as an input to another function, or return a function from a function, you can have arrays of functions, you can create functions on the fly.
Passing functions as an input to another function
| |
Returning a function from a function
| |
This is of course of limited value. Here is a really useful one:
| |
You can numerically solve the equation :
using this function as: iter(cos,100)
| |
Special concepts
source():
source()is used to execute R code stored in an external script or file. It is commonly used to read and execute a script file and load its functions and variables into the R environment.
| |
browser():
browser()is used for interactive debugging in R. When placed within a function, it suspends the execution of the function, and the user can interactively inspect variables, evaluate expressions, and step through code.- It is often used for setting breakpoints and examining the program’s state during debugging.
| |
invisible():
invisible()is a function used to make the return value of a function “invisible” to the console. It is often used when you want a function to return a value, but you don’t want that value to be printed to the console.- This can be useful when a function is meant to produce a side effect or when you don’t want the function’s result to clutter the console output.
Here is the code:
| |
Closures:
- A closure in R is a function that “closes over” its environment, meaning it retains access to the variables in the environment where it was created, even after that environment goes out of scope.
- Closures are created when a function is defined within another function and references variables from the outer function’s environment.
- Closures are useful for creating functions with persistent state.
| |
Apply:
apply() function is used to apply a given function to the rows or columns of a matrix or array.
apply(X, MARGIN, FUN, ...)X: The data matrix or array on which you want to apply the function.MARGIN: An integer vector specifying whether the function should be applied over rows (MARGIN = 1) or columns (MARGIN = 2).FUN: The function to be applied to each row or column....: Additional arguments to be passed to the function specified inFUN.
Stats
variance/covariance/correlation
| |
| x | a numeric vector, matrix or data frame. |
|---|---|
| y | NULL (default) or a vector, matrix or data frame with compatible dimensions to x. The default is equivalent to y = x (but more efficient). |
| na.rm | logical. Should missing values be removed? |
| use | an optional character string giving a method for computing covariances in the presence of missing values. This must be (an abbreviation of) one of the strings “everything”, “all.obs”, “complete.obs”, “na.or.complete”, or “pairwise.complete.obs”. |
var, cov and cor compute the variance of x and the covariance or correlation of x and y if these are vectors. If x and y are matrices then the covariances (or correlations) between the columns of x and the columns of y are computed.
Sure! Here’s the code rewritten with detailed explanations suitable for beginners:
R-squared (R²) and Multiple Correlation
R-squared (R²), also known as the coefficient of determination, is a statistical measure that represents the proportion of the variance for a dependent variable that’s explained by independent variables in a regression model. Essentially, it indicates how well the independent variables predict the dependent variable.
- Formula:
Where:
- Sum of Squared Residuals (SSR): The sum of the squared differences between the observed values and the predicted values.
- Total Sum of Squares (TSS): The sum of the squared differences between the observed values and the mean of the observed values.
Interpretation:
- An R² of 1 indicates that the regression predictions perfectly fit the data.
- An R² of 0 indicates that the model does not explain any of the variability of the response data around its mean.
- Values between 0 and 1 indicate the extent to which the independent variables explain the variability of the dependent variable.
Multiple correlation refers to the correlation between one dependent variable and a set of independent variables. It assesses the strength of the relationship between the dependent variable and the combination of independent variables.
- Formula:
Where R² is the coefficient of determination from a multiple regression analysis.
Interpretation:
- The multiple correlation coefficient ( R ) ranges from 0 to 1.
- An R close to 1 indicates a strong relationship between the dependent variable and the set of independent variables.
- An R close to 0 indicates a weak relationship.
| |
How to interpret the results?
summary(tmp)
Residual standard error: 0.015 on 17 degrees of freedom (model fits better - lower residual variation)
Multiple R-squared: 0.995 (more R squared - model explains more variance)
Adjusted R-squared: 0.994
Factors in R
In R, factors are variables used to represent categorical data, storing both the values and their corresponding category labels. This is particularly useful for statistical modeling and data analysis, as factors ensure that categorical variables are treated appropriately.
Creating Factors in R:
To create a factor in R, you can use the factor() function. For example:
| |
In this example, treatment_factor will store the categorical data along with its distinct levels: “Control”, “Treatment1”, and “Treatment2”.
Using Factors in Linear Models:
When performing linear regression in R, incorporating factors allows the model to account for categorical predictors. The lm() function automatically recognizes factor variables and handles them accordingly.
Consider the following linear model:
| |
In this model:
post: The dependent variable (response).pre: An independent variable (predictor).treatment: A categorical predictor indicating different treatment groups.leprosy: The dataset containing these variables.
By using factor(treatment), R treats the treatment variable as categorical, allowing the model to estimate separate effects for each treatment group compared to a reference group. This approach is essential when the predictor variable represents distinct categories rather than continuous data.
Partial Correlation
Partial correlation measures the strength and direction of the relationship between two continuous variables while controlling for the effect of one or more other continuous variables. It helps to understand the direct relationship between two variables by removing the influence of other variables.
| |
An interesting feature of R is that allows arbitrary meta-information to be associated with any variable. These are called attributes.
We want to associate an attribute date with it. Then we shall use attr(x,’date’) = "June 6, 2011"
You may find the attributes associated with an object using the attributes function:
attributes(x) #Looking up all attributes.
attr(x,’date’) #Looking up a specific attribute.
Certain attributes are special and are used internally by R. One such is the dim attribute that stores the dimension of an array.
Simulation and Statistic
You have a random experiment, and you want to do some math with its outcome, and save the result. You repeat this a large number of times (say ) to obtain a long vector of results. Finally, look at the results through the lens of statistical regularity, i.e., draw histogram or compute mean, or whatever. The basic structure is:
| |
You have a random experiment, and you want to do some math with its outcome, and save the result. You repeat this a large number of times (say ) to obtain a long vector of results. Finally look at the results through the lens of statistical regularity, i.e., draw histogram or compute mean, or whatever. The basic structure is
| |
EXAMPLE:-
We may toss a coin 100 times in each pass, and save the number of heads obtained. If we repeat this 10
times, then we shall end up with a vector nhead of length 10, where nhead[i] stores the number of heads in the -
th batch of 100 tosses. Here is a code to do this:
| |
Approximating something using simulation
Often we want to determine some quantity related to a statistical model. Since any statistical model involves some random experiment, it may become hard to determine the quantity. Indeed, probability theory is designed mainly to cope with this problem. But even the most sophisticated probability theory often proves inadequate to deal with even simple statistical models. So we need an alternative , Simulation is just that alternative, it is very general in its applicability, pretty routine to apply (once you get the hang of it). It provides only approximate answers, but then so does probability theory (it approximates using limit).
Finding standard error of something
Suppose that you have some statistical model, and you have somehow come up with some statistic based on them, i.e. some known function of the data. It could be something as simple as the mean of the data, or as complicated as the prediction to tomorrow’s rainfall based on past 10 years’ data. The function could be something mathematical, or a complicated program. But the function is known, i.e., given the data you have some means to compute it. Now you want to have an idea about the variability of its values (since its input is random). In this situation we compute the standard error of the statistic: it is simply the standard deviation of the statistic.
The general R code skeleton to compute standard error is:
| |
Finding sampling distribution of something
If you want a broader perspective, then you might like to investigate the sampling distribution of your statistic, i.e., the probability distribution of the statistic, instead of just its standard error.
The R skeleton remains the same, except that the last line is changed to
hist(myStatistic,prob=T)
Finding probability of something
If you want probability of some event (like the probability that your statistic is less than 3.4), then you just simulate lots of values of your statistic, and find the proportion of cases your event has occured. If the event is “statistic is less than 3.4”, then simply replace the last line of our R skeleton with
mean(myStatistic < 3.4)
Finding cut-off values
Often you want to find cut-off thresholds for your statistic, e.g., an upper bound that it crosses with 5% probability. We may find this using simulation by first sorting the simulated values of the statistic in ascending order, and then picking the cut-off where the top 5% values start. The R function quantile does this for you:
quantile(myStatistic,0.95)
This finds a cut-off such that 95% of the statistic values are below it, and 5% are above it. Now, you may easily guess that this rather vague description has problems: what if we have no such cut-off, more than one such cut-off? There are different ways to solve this problem. If you look up the help of the quantile function, you will find a input called type that chooses the specific algorithm to be used. However, if you do the simulation a large number of times (i.e., after statistical regularity has set in), all the methods give you essentially the same answer for the continuous distribution. So you do not need to bother much about the exact algorithm being used. Generally, we refrain from using the quantile function for the discrete case
Fitting Distribution
Linear regression
lm() function is used to fit linear regression models. Linear regression is a statistical method used to model the relationship between a dependent variable and one or more independent variables by fitting a linear equation to the observed data.
| |
| |
In R, the set.seed() function is used to set the seed for generating random numbers using various random number generators. This is important when you want to ensure that your code generates the same random numbers each time it’s run, making your results reproducible.
| |
abline(lm(y ~ x), col = "purple") adds a linear regression line to the plot using the lm() function to fit a linear model to the data. It’s colored purple.
names(lm(x ~ y)) would return the names of the coefficients or parameters estimated by the linear regression model created using the lm() function. These names correspond to the coefficients that represent the intercept and the slope(s) for the independent variable(s) in the regression model.
In R, when you calculate the variance using the var() function, it may not always match the manual calculation of the variance due to differences in the formula used and potential differences in data handling. The discrepancy could be attributed to the following factors:
Degree of Freedom (Bessel’s correction):
- By default, the
var()function in R calculates the sample variance with Bessel’s correction, which divides the sum of squared differences by (n - 1), where ’n’ is the number of data points. This correction is used when working with sample data to provide an unbiased estimate of the population variance. - If you manually calculate the variance without applying Bessel’s correction by dividing by ’n’ instead of ’n - 1’, the result will differ.
- To avoid try var(x) * (n-1)/n, where n in length(n)
Univariate distributions
fitdistr(x, densfun, start, ...) # Maximum-likelihood fitting of univariate distributions, allowing parameters to be held fixed if desired
| x | A numeric vector of length at least one containing only finite values. |
|---|---|
| densfun | Either a character string or a function returning a density evaluated at its first argument. Distributions “beta”, “cauchy”, “chi-squared”, “exponential”, “gamma”, “geometric”, “log-normal”, “lognormal”, “logistic”, “negative binomial”, “normal”, “Poisson”, “t” and “weibull” are recognised, case being ignored. |
| |
Distributions
THE NORMAL DISTRIBUTION
| |
Density, distribution function, quantile function and random generation for the normal distribution with mean equal to mean and standard deviation equal to sd. If mean or sd are not specified they assume the default values of 0 and 1, respectively.
| x, q | vector of quantiles. |
|---|---|
| p | vector of probabilities. |
| n | number of observations. If length(n) > 1, the length is taken to be the number required. |
| mean | vector of means. |
| sd | vector of standard deviations. |
Multivariate Normal Density and Random Deviates
Description
These functions provide the density function and a random number generator for the multivariate normal distribution with mean equal to mean and covariance matrix sigma.
Usage
| |
Arguments
| x | vector or matrix of quantiles. When x is a matrix, each row is taken to be a quantile and columns correspond to the number of dimensions, p. |
|---|---|
| n | number of observations. |
| mean | mean vector, default is rep(0, length = ncol(x)). In ldmvnorm or sldmvnorm, mean is a matrix with observation-specific means arranged in columns. |
| sigma | covariance matrix, default is diag(ncol(x)). |
| log | logical; if TRUE, densities d are given as log(d). |
Details
dmvnorm computes the density function of the multivariate normal specified by mean and the covariance matrix sigma.
rmvnorm generates multivariate normal variables.
EXAMPLE:-
dat = rmvnorm(100000,c(0,0),sigma=matrix(c(2,1,1,2),2,2)) # This code generates 100,000 random samples from a bivariate normal distribution with mean (0, 0) and a covariance matrix: Σ=(2,1;1,2)
The (non-central) Chi-Squared Distribution
Description
Density, distribution function, quantile function and random generation for the chi-squared (χ2) distribution with df degrees of freedom and optional non-centrality parameter ncp.
Usage
| |
Arguments
| x, q | vector of quantiles. |
|---|---|
| p | vector of probabilities. |
| n | number of observations. If length(n) > 1, the length is taken to be the number required. |
| df | degrees of freedom (non-negative, but can be non-integer). |
| log, log.p | logical; if TRUE, probabilities p are given as log(p). |
Values
dchisq gives the density, pchisq gives the distribution function, qchisq gives the quantile function, and rchisq generates random deviates.
Invalid arguments will result in return value NaN, with a warning.
The length of the result is determined by n for rchisq, and is the maximum of the lengths of the numerical arguments for the other functions.
The numerical arguments other than n are recycled to the length of the result. Only the first elements of the logical arguments are used.
Examples
| |
The Poisson Distribution
Density, distribution function, quantile function and random generation for the Poisson distribution with parameter lambda . dpois gives the (log) density, ppois gives the (log) distribution function, qpois gives the quantile function, and rpois generates random deviates.
| |
| x | vector of (non-negative integer) quantiles. |
|---|---|
| q | vector of quantiles. |
| p | vector of probabilities. |
| n | number of random values to return. |
| lambda | vector of (non-negative) means. |
| log, log.p | logical; if TRUE, probabilities p are given as log(p). |
| lower.tail | logical; if TRUE (default), probabilities are P[X≤x], otherwise, P[X>x]. |
Some function used frequently
sample(x, size=n, replace = FALSE, prob = NULL)#sampletakes a sample (random) of the specified size from the elements ofxusing either with (replace = T) or without replacement with probability weights given as a vector, DEFAULT:- prob=rep(1,n) # that is equally likelyexample :-
sample(c("Heads", "Tail"),100,replace=T)runif(n, min = a, max = b)#generates random deviates/numbers (uniform distribution) between a and b (default 0 and 1)
Contingency tables
Table
table(vector) # uses cross-classifying factors to build a contingency table of the counts at each combination of factor levels.
| |
as.table and is.table coerce to and test for contingency table, respectively.
XTABS
Create a contingency table (optionally a sparse matrix) from cross-classifying factors, usually contained in a data frame, using a formula interface.
| |
| parameter | details |
|---|---|
| formula | a formula object with the cross-classifying variables (separated by +) on the right hand side (or an object which can be coerced to a formula). |
| data | an optional matrix or data frame containing the variables in the formula formula. By default the variables are taken from environment(formula). |
| subset | an optional vector specifying a subset of observations to be used. |
examples: -
| |
| |
margin.table
For a contingency table in array form, compute the sum of table entries for a given margin or set of margins.
margin.table(x, margin = NULL)
| parameter | details |
|---|---|
| x | an array |
| margin | a vector giving the margins to compute sums for. E.g., for a matrix 1 indicates rows, 2 indicates columns, c(1, 2) indicates rows and columns. When x has named dimnames, it can be a character vector selecting dimension names. |
example:-
| |
margin.table(ft, 1): This function calculates the marginal totals for the rows of the tableft. The argument1indicates that the operation is applied to the rows. The result is a vector containing the sum of counts for each level of the first margin (rows).(vicTot=...): The result of themargin.tableoperation is assigned to the variablevicTotusing the assignment operator=. This is a convenient way of both calculating the marginal totals and storing the result in a variable in a single line.
Data Handling
DATA CLEANING
| |
Handle Missing Values in Objects
In R, handling missing values is crucial for data analysis. Several functions are available to manage these missing values in different ways, depending on how you want to treat them during analysis. Here’s an explanation of some common functions:
na.fail- This function ensures that the data has no missing values. If any missing values are found, it stops the function and returns an error.
- Usage:
na.fail(object, ...) - Example:This will produce an error because there are missing values in
1 2data <- data.frame(x = c(1, 2, NA), y = c(4, NA, 6)) na.fail(data)data.
na.omit- This function removes rows with any missing values from the data frame.
- Usage:
na.omit(object, ...) - Example:This will return a data frame with rows containing missing values removed:
1 2 3data <- data.frame(x = c(1, 2, NA), y = c(4, NA, 6)) clean_data <- na.omit(data) print(clean_data)1 2x y 1 4
na.exclude- Similar to
na.omit, this function removes rows with missing values but keeps the positions of NA values in residuals and predictions for time series analysis. - Usage:
na.exclude(object, ...) - Example:This will also return a data frame with rows containing missing values removed. The main difference is how residuals and predictions are handled in models.
1 2 3data <- data.frame(x = c(1, 2, NA), y = c(4, NA, 6)) clean_data <- na.exclude(data) print(clean_data)
- Similar to
na.pass- This function does nothing to the data and allows all NA values to pass through.
- Usage:
na.pass(object, ...) - Example:This will return the original data frame, unchanged:
1 2 3data <- data.frame(x = c(1, 2, NA), y = c(4, NA, 6)) clean_data <- na.pass(data) print(clean_data)1 2 3 4x y 1 4 2 NA NA 6
Data Manipulation and analysis
We will understand this with example of iris dataset:
Overview
data() # loads all and gives the names of all inbuilt databases in R
data(iris) #for specific loading
iris
This famous (Fisher’s or Anderson’s) iris data set gives the measurements in centimeters of the variables sepal length and width and petal length and width, respectively, for 50 flowers from each of 3 species of iris. The species are Iris setosa, versicolor, and virginica.
dim(iris) # 150 5
names(iris) # gives names of the headings of the data set
names() function is used to access or modify the names of elements within various data structures such as vectors, lists, data frames, and arrays
| |
Insights
plot(iris) #The plot() function, when used with the iris dataset, by default, creates a matrix of scatter plots, where each feature is plotted against every other feature. In total, it produces a 5*5 grid of scatter plots (5 containing only names(headings) which act as the coordinates for the scatterplots), showing the relationships between the different features of the iris flowers.
iris[,1] = 0 # all rows of first column of iris = 0
rm(iris) # will remove the “iris” dataset from your current R session. After executing this command, you won’t be able to access the “iris” dataset or its variables until you load it again or restart R. It’s essentially a way to free up memory and remove unnecessary objects from your workspace when you no longer need them.
The tail() function is used to display the last few rows of a dataset. By default, it shows the last 6 rows of the specified dataset
head(): The head() function is used to display the first few rows of a dataset. By default, it shows the first 6 rows of the specified dataset.
#The key difference is which subset of the data you are viewing. The head() command displays the beginning of the subset, and the tail() command displays the end of the subset. The content of the displayed data will differ because you’re looking at different rows within the dataset. If the rows within the subset are unique and not repeated in the dataset, these commands will display different rows of data.
| |
iris[50:56,c("Petal.Length","Sepal.Width")] #print row 50 to 56 of petal length and sepal width
iris[c(45,78,98),c("sw","sl")] names(iris) = c("sl","sw","pl","pw","spec")#changes names of the headers
iris[c(45,78,98),c("sw","sl")]#gives respected rows of respected columns after changing the names
sw #Error: object ‘sw’ not found
attach(iris) #The database is attached to the R search path. This means that the database is searched by R when evaluating a variable, so objects in the database can be accessed by simply giving their names., now you can access columns by the changed names
iris[ sw>4 , ] #print all columns where spec>4
iris[ sw>4 , 'spec'] #print spec columns where spec>4
detach(iris) #Detach a database, i.e., remove it from the search() path of available R objects. Usually this is either a data.frame which has been attached or a package which was attached by library.
pairs(x, ...)
x: the coordinates of points given as numeric columns of a matrix or data frame. Logical and factor columns are converted to numeric in the same way that data.matrix does.
| |
Visualisation
library(rgl) #3D visualization device system ; Description : 3D real-time rendering system. ;Details: RGL is a 3D real-time rendering system for R. Multiple windows are managed at a time.
options(rgl.printRglwidget = TRUE) # TRUE means that R will try to display interactive 3D graphics created with the “rgl” library in your current environment. This can be useful for visualizing 3D plots interactively.
plot3d(x, ...)
plot3d(x, y, z, xlab, ylab, zlab, type = "p", col, size, lwd, radius, add = FALSE, aspect = !add, xlim = NULL, ylim = NULL, zlim = NULL, forceClipregion = FALSE, decorate = !add, ...)
| parameters | details |
|---|---|
| x, y, z | vectors of points to be plotted. |
| xlab, ylab, zlab | labels for the coordinates. |
| type | For the default method, a single character indicating the type of item to plot. Supported types are: ‘p’ for points, ’s’ for spheres, ’l’ for lines, ‘h’ for line segments from z = 0, and ’n’ for nothing. |
| col | the color to be used for plotted items. |
| size | the size for plotted points. |
| lwd | the line width for plotted items. |
| xlim, ylim, zlim | If not NULL, set clipping limits for the plot. |
| |
library(aplpack) #This line loads the “aplpack” library, which provides various functions for creating different types of plots, including “faces” plots.
faces(iris[,-5]) #This command generates a “faces” plot using the Iris dataset with the fifth column (species) excluded. The “faces” plot is a visualization technique that represents multivariate data in a grid of small faces, where each face corresponds to an observation in the dataset. Different facial expressions and features are used to encode the values of the variables.
faces(iris[sample(150,20),-5]) #This command creates another “faces” plot, but this time it’s based on a random sample of 20 observations from the Iris dataset, excluding the fifth column (species). This will give you a “faces” plot for a subset of the Iris data.
Other datasets
Similarly there are are more in-built datasets in R you can work with:
sunspot.year#Yearly numbers of sunspots from 1700 to 1988 (rounded to one digit).The univariate time series sunspot.year contains 289 observations, and is of class “ts”plot(sunspot.year, ty='l')#sunspot.year: This is the data you want to (line) plot. It appears to be a time series of sunspot observations over years. Each data point likely represents the number of sunspots observed in a given year.quakes#The data set give the locations of 1000 seismic events of MB > 4.0. The events occurred in a cube near Fiji since 1964. A data frame with 1000 observations on 5 variables.- [,1] lat numeric Latitude of event
- [,2] long numeric Longitude
- [,3] depth numeric Depth (km)
- [,4] mag numeric Richter Magnitude
- [,5] stations numeric Number of stations reporting
plot(quakes[,-5])#create a scatterplot of earthquake locations, where the longitude is on the x-axis, latitude is on the y-axis, and we’ve added labels to the axes and a title to the plot.
PLOT, LOCATOR and POINTS
plot(1:10, 1:10) # This just creates scatterplot with the points ( for Now issue the command)
locator(n = 512, type = "n", ...) #Reads the position of the graphics cursor when the (first) mouse button is pressed.
If you use locator() without specifying the number of clicks, then it allows any number of clicks (you have to terminate by right-clicking).
| parameter | details |
|---|---|
| n | the maximum number of points to locate. Valid values start at 1. |
| type | One of “n”, “p”, “l” or “o”. If “p” or “o” the points are plotted; if “l” or “o” they are joined by lines. |
example:-
locator(1)#The prompt will go away, indicating that the program is running waiting for input from you. Visit the plot window, take your mouse over the plot region. You will notice the mouse cursor changing to a crosshair. Click at some point.Immediately the R prompt will return and coordinates (wrt the axis used in the plot) of the click point will be shown on screen. The 1 passed to locator tells it to return after a single click.repeat {locator(2,ty="l"}#creates an interactive loop where the user is prompted to click on two points along lines on a plot. The loop will continue indefinitely until the user decides to stop it manually (e.g., by closing the plot window), as there’s no explicit condition to break the loop in the provided code.p= locator(3)p$x#All the x valuesp$y#All the y valuesp$x[1]#x component of the first clickp$y[1]#y component of the first click
The function points adds points to an existing plot. Points outside the current plot region are silently ignored.
example:-
| |
Data Visualisation
curve
The curve function in R is used to plot a curve corresponding to a function over a specified range of x values. This function is very useful for visualizing mathematical functions and probability distributions.
curve(dchisq(x,df=10),xlim=c(0,50),col='black',lwd=5) # generates a plot of the chi-squared probability density function with 10 degrees of freedom, evaluated over the x-axis range from 0 to 50, and displays it as a thick (line width of 5), black curve.
PLOT
plot(x, y = NULL, type = "p", xlim = NULL, ylim = NULL, main = NULL, xlab = NULL, ylab = NULL, axes = TRUE) # Draws a scatter plot with decorations such as axes and titles in the active graphics window.
| parameter | details |
|---|---|
| x, y | the x and y arguments provide the x and y coordinates for the plot. Any reasonable way of defining the coordinates is acceptable. If supplied separately, they must be of the same length. |
| type (ty) | 1-character string giving the type of plot desired. The following values are possible: “p” for points, “l” for lines, “b” for both points and lines, “c” for empty points joined by lines, “o” for overplotted points and lines, “s” and “S” for stair steps and “h” for histogram-like vertical lines. Finally, “n” does not produce any points or lines. |
| xlim | the x limits (x1, x2) of the plot. Note that x1 > x2 is allowed and leads to a ‘reversed axis’. The default value is NULL. |
| ylim | the y limits of the plot. |
| main | a main title for the plot, see also title. |
| sub | a subtitle for the plot. |
| xlab | a label for the x axis, defaults to a description of x. |
| ylab | a label for the y axis, defaults to a description of y. |
| col | The colors for lines and points. Multiple colors can be specified so that each point can be given its own color. If there are fewer colors than points they are recycled in the standard fashion. Lines will all be plotted in the first colour specified; e.g., col=rgb(0,0,1,1/4) different colour combinations can be made, 1/4 signifies the opacity of chosen color |
| bg | a vector of background colors for open plot symbols, see points. Note: this is not the same setting as par(“bg”) |
| pch | a vector of plotting characters or symbols: see points. |
| lty | a vector of line types, see par |
| lwd | a vector of line widths, see par. |
examples :-
plot(h1, col=rgb(0,0,1,1/4), xlim=c(-40000,40000), xlab="Histogram")
plot(x[,1],qnorm(p),pch = 20) # pch is for different types of dots available with different integers
plot(1:100,cumsum(sample(2,100,replace=T)==2)/1:100)) # guess the output then try it yourself
Let’s see some practical examples:-
Example 1:
| |
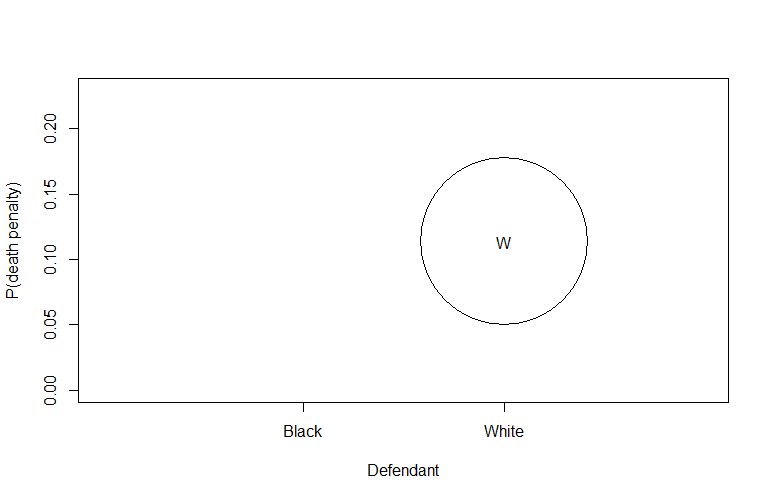
Example 2:
| |
This line initializes a plot with specific settings:
1is a placeholder for the x-axis values (in this case, there’s no actual data on the x-axis).ylim=range(dp)sets the y-axis limits based on the range of values in thedpmatrix.xlim=c(0,3)sets the x-axis limits.xlab=''andylab=''specify empty labels for the x and y axes.ty='n'specifies that no points should be plotted initially.xaxt='n'prevents drawing the x-axis.
| |
BARPLOT
barplot(height, width = 1, space = NULL, names.arg = NULL, col = NULL, main = NULL, xlab = NULL, ylab = NULL, xlim = NULL, ylim = NULL, axes = TRUE, axisnames = TRUE) # Creates a bar plot with vertical or horizontal bars.
| parameter | details |
|---|---|
| height | either a vector or matrix of values describing the bars which make up the plot. |
| width | optional vector of bar widths. Re-cycled to length the number of bars drawn. Specifying a single value will have no visible effect unless xlim is specified. |
| names.arg | a vector of names to be plotted below each bar or group of bars. If this argument is omitted, then the names are taken from the names attribute of height if this is a vector, or the column names if it is a matrix. |
| col | a vector of colors for the bars or bar components. By default, “grey” is used if height is a vector, and a gamma-corrected grey palette if height is a matrix. |
| border | the color to be used for the border of the bars. Use border = NA to omit borders. If there are shading lines, border = TRUE means use the same colour for the border as for the shading lines. |
| main, sub | main title and subtitle for the plot. |
| xlab | a label for the x axis. |
| ylab | a label for the y axis. |
| xlim | limits for the x axis. |
| ylim | limits for the y axis. |
| axisnames | logical. If TRUE, and if there are names.arg (see above), the other axis is drawn (with lty = 0) and labeled. |
try examples: -
barplot(table(sample(c("Heads", "Tail"),100,replace=T)))barplot(table(rpois(1000,lambda=5))/1000)barplot(table(rbinom(100,10,0.3))/100)
Legend
This function can be used to add legends to plots. Note that a call to the function locator(1) can be used in place of the x and y arguments.
legend(x, y = NULL, legend, fill = NULL, col = par("col"), border = "black", lty, lwd, pch, angle = 45, density = NULL)
| parameter | details |
|---|---|
| x, y | the x and y co-ordinates to be used to position the legend. They can be specified by keyword or in any way which is accepted by xy.coords: See ‘Details’. |
| legend | a character or expression vector of length \ge 1≥1 to appear in the legend. Other objects will be coerced by as.graphicsAnnot. |
| fill | if specified, this argument will cause boxes filled with the specified colors (or shaded in the specified colors) to appear beside the legend text. |
| col | the color of points or lines appearing in the legend. |
| border | the border color for the boxes (used only if fill is specified). |
| lty, lwd | the line types and widths for lines appearing in the legend. One of these two must be specified for line drawing. |
| pch | the plotting symbols appearing in the legend, as numeric vector or a vector of 1-character strings (see points). Unlike points, this can all be specified as a single multi-character string. Must be specified for symbol drawing. |
| angle | angle of shading lines. |
| density | the density of shading lines, if numeric and positive. If NULL or negative or NA color filling is assumed. |
| bty | the type of box to be drawn around the legend. The allowed values are “o” (the default) and “n”. |
| bg | the background color for the legend box. (Note that this is only used if bty != “n”.) |
Lines
lines(x, y = NULL, type = "l", ...) #A generic function taking coordinates given in various ways and joining the corresponding points with line segment,
| parameter | details |
|---|---|
| x, y | coordinate vectors of points to join. |
| type | character indicating the type of plotting |
example :-
lines(ntoss,cumsum(sample(2,100,replace=T)-1)/ntoss,col = 'red')
lines(c(5,20),c(5,20)) # line from (5,5) to (20,20)
NOTE:- generally “lines” plots on the pre plotted graph only without specifying any add = T
abline
abline() function is used to add straight lines to a plot, typically a scatterplot or other types of plots.
abline(h = 4, col = "red")adds a horizontal line aty = 4with a red color.abline(v = 3, col = "blue")adds a vertical line atx = 3with a blue color.abline(a = 1, b = 1, col = "green")adds a diagonal line with a slope of 1 and an intercept of 1, colored green.abline(0,1)adds a diagonal line with slope 1 and intercept 0.
Histograms
1-D histograms
hist(x, breaks = "Sturges", freq = NULL, probability = !freq, include.lowest = TRUE, col = "lightgray", border = NULL, main = paste("Histogram of" , xname), xlim = range(breaks), ylim = NULL, xlab = xname, ylab, axes = TRUE, plot = TRUE, labels = FALSE, nclass = NULL)
| parameter | details |
|---|---|
| x | a vector of values for which the histogram is desired. |
| breaks | one of: 1. a vector giving the breakpoints between histogram cells, 2. a function to compute the vector of breakpoints, 3. a single number giving the number of cells for the histogram, 4. a character string naming an algorithm to compute the number of cells, 5. a function to compute the number of cells. |
| freq | logical; if TRUE, the histogram graphic is a representation of frequencies, the counts component of the result; if FALSE, probability densities, component density, are plotted (so that the histogram has a total area of one). Defaults to TRUE if and only if breaks are equidistant (and probability is not specified). |
| probability | an alias for !freq, for S compatibility. (when True gives probability density). The formula for probability density (PD) in a bin is: PD = (Count in Bin) / (Bin Width * Total Number of Data Points) where; • Count in Bin: The raw count of data points in the bin. • Bin Width: The width of the bin along the x-axis. • Total Number of Data Points: The total number of data points in your dataset. |
| col | a colour to be used to fill the bars. |
| border | the color of the border around the bars. The default is to use the standard foreground color. |
| main, xlab, ylab | main title and axis labels |
| xlim, ylim | the range of x and y values with sensible defaults. Note that xlim is not used to define the histogram (breaks), but only for plotting (when plot = TRUE). |
| axes | logical. If TRUE (default), axes are draw if the plot is drawn. |
| plot | logical. If TRUE (default), a histogram is plotted, otherwise a list of breaks and counts is returned. In the latter case, a warning is used if (typically graphical) arguments are specified that only apply to the plot = TRUE case. |
examples :-
hist(x,n=785) # 785 bars
hist(cumsum(a)/b, xlim=c(0,2), prob = T, breaks = c(1)) # x axis from 0 to 2, breaks at 1
2-D histograms
hist2d(x,y=NULL, nbins=200, na.rm=TRUE, show=TRUE, col=c("black", heat.colors(12)), xlab, ylab, ... ) # Compute and plot a 2-dimensional histogram.
Arguments
| parameter | details |
|---|---|
| x | either a vector containing the x coordinates or a matrix with 2 columns. |
| y | a vector contianing the y coordinates, not required if ‘x’ is matrix |
| nbins | number of bins in each dimension. May be a scalar or a 2 element vector. Defaults to 200. |
| show | Indicates whether the histogram be displayed using image once it has been computed. Defaults to TRUE. |
| col | Colors for the histogram. Defaults to “black” for bins containing no elements, a set of 16 heat colors for other bins. |
| xlab, ylab | (Optional) x and y axis labels |
Details
This fucntion creates a 2-dimensional histogram by cutting the x and y dimensions into nbins sections. A 2-dimensional matrix is then constucted which holds the counts of the number of observed (x,y) pairs that fall into each bin. If show=TRUE, this matrix is then then passed to image for display.
2D histogram provides a way to visualize the joint distribution of two variables by dividing the data space into bins and counting the number of observations in each bin. It is particularly useful for identifying patterns and structures in the relationship between the variables.
Examples:-
| |
3-D histograms
Description
hist3D (x = seq(0, 1, length.out = nrow(z)), y = seq(0, 1, length.out = ncol(z)), z) # generates 3-D histograms.
| parameter | details |
|---|---|
| z | Matrix (2-D) containing the values to be plotted as a persp plot. |
| x, y | Vectors or matrices with x and y values. If a vector, x should be of length equal to nrow(z) and y should be equal to ncol(z). If a matrix (only for persp3D), x and y should have the same dimension as z. |
| col | Color palette to be used for the colvar variable. |
| NAcol | Color to be used for NA values of colvar; default is “white”. |
| breaks | a set of finite numeric breakpoints for the colors; must have one more breakpoint than color and be in increasing order. Unsorted vectors will be sorted, with a warning. |
| colkey | A logical, NULL (default), or a list with parameters for the color key (legend). |
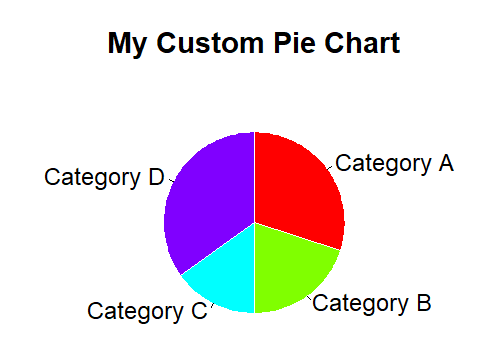
pie()
pie(x, labels = names(x), edges = 200, radius = 0.8, clockwise = FALSE, init.angle = if(clockwise) 90 else 0, density = NULL, angle = 45, col = NULL, border = NULL, lty = NULL, main = NULL, ...) # Draw a pie chart.
| parameter | details |
|---|---|
| x | a vector of non-negative numerical quantities. The values in x are displayed as the areas of pie slices. |
| labels | one or more expressions or character strings giving names for the slices. |
| edges | the circular outline of the pie is approximated by a polygon with this many edges. |
| radius | the pie is drawn centered in a square box whose sides range from −1−1 to 11. If the character strings labeling the slices are long it may be necessary to use a smaller radius. |
| clockwise | logical indicating if slices are drawn clockwise or counter clockwise (i.e., mathematically positive direction), the latter is default. |
| init.angle | number specifying the starting angle (in degrees) for the slices. Defaults to 0 (i.e., ‘3 o’clock’) unless clockwise is true where init.angle defaults to 90 (degrees), (i.e., ‘12 o’clock’). |
| density | the density of shading lines, in lines per inch. The default value of NULL means that no shading lines are drawn. Non-positive values of density also inhibit the drawing of shading lines. |
| angle | the slope of shading lines, given as an angle in degrees (counter-clockwise). |
| col | a vector of colors to be used in filling or shading the slices. If missing a set of 6 pastel colours is used, unless density is specified when par(“fg”) is used. |
Example:-
| |
Files Handling
READTABLE & ReadCSV
read.table(file, header = FALSE, sep = "", row.names, col.names, nrows = -1, skip = 0)
read.csv(file, header = TRUE, sep = ",")
| parameter | details |
|---|---|
| file | the name of the file which the data are to be read from. Each row of the table appears as one line of the file. If it does not contain an absolute path, the file name is relative to the current working directory |
| header | a logical value indicating whether the file contains the names of the variables as its first line. If missing, the value is determined from the file format: header is set to TRUE if and only if the first row contains one fewer field than the number of columns. If head = T, then the first row is made to be the header rather than V(i)’s and now it can be accessed that way too that is using (table_name)$(column_name) |
| sep | the field separator character. Values on each line of the file are separated by this character. If sep = "” (the default for read.table) the separator is ‘white space’, that is one or more spaces, tabs, newlines or carriage returns. |
| row.names | a vector of row names. This can be a vector giving the actual row names, or a single number giving the column of the table which contains the row names, or character string giving the name of the table column containing the row names. |
| col.names | a vector of optional names for the variables. The default is to use “V” followed by the column number. |
| nrows | integer: the maximum number of rows to read in. Negative and other invalid values are ignored. |
| skip | integer: the number of lines of the data file to skip before beginning to read data. |
Example:-
| |
The $ operator can be used to select a variable/column, assign new values to a variable/column, or add a new variable/column in an R object. This R operator can be used on e.g., lists, and dataframes.
For example, if we want to print the values in the column “A” in the dataframe called “dollar,” we can use the following code:
print(dollar$A).First, using the double brackets enables us to select multiple columns, e.g., whereas the $ operator only enables us to select one column.
so x[,1] = x$V1 or like x[,2] = x$col2
Text Files
File handling in R involves two main tasks: writing data to files and reading data from files. Here’s an overview of how to handle these tasks:
Writing to Files:
- The primary function for outputting data in R is
print, which displays data in a formatted way on the screen. To redirect this output to a file, use thesinkfunction.1 2 3 4x = matrix(1:12, 3, 4) # Creating an example matrix sink("myfile.txt") # Redirecting output to a file summary(x) # Output is redirected to 'myfile.txt' sink() # Reverting output back to the console
- The primary function for outputting data in R is
Custom File Output:
- For finer control over file output, the
catfunction can be used.1 2 3y = 20 cat("The answer is", y, "\n", file="myfile.txt") # Writing a string to 'myfile.txt' cat("This is a second line\n", file="myfile.txt", append=TRUE) # Appending another line
- For finer control over file output, the
Writing Objects to Files:
- To save the content of an R object in a human-readable format, the
writefunction is useful.1 2x = matrix(1:12, 3, 4) # Creating an example matrix write(x, ncol=4, file="myfile.txt") # Writing matrix 'x' to 'myfile.txt'
- To save the content of an R object in a human-readable format, the
Saving and Loading Objects:
- If the goal is to save an R object for future use, the
saveandloadfunctions are the most straightforward approach.1 2 3save(x, file="abc.RData") # Saving the object 'x' to 'abc.RData' rm(x) # Removing the object 'x' load("abc.RData") # Loading the object 'x' from 'abc.RData'
- If the goal is to save an R object for future use, the
Audio handling
install.packages(tuneR)
library(tuneR) #importing tuneR package
tuneR consists of several functions to work with and to analyze Wave files. In the following examples, some of the functions to generate some data (such as sine), to read and write Wave files (readWave, writeWave), to represent or construct (multi channel) Wave files (Wave, WaveMC), to transform Wave objects (bind, channel, downsample, extractWave, mono, stereo), and to play Wave objects are used.
s = readWave('recording.wav') #reading .wav file
str(s) # Compactly display the internal structure of an R object, a diagnostic function and an alternative to summary (here, audio file)
example: -
| |
s@left or s@right #extracting left or right part of our audio
object@nameExtract or replace the contents of a slot or property name given of an object.
plotting s@left using diffrent functions
hist(s@left,prob=T,breaks=seq(-15000,15000,1000))hist(s@left,prob=T,breaks=seq(min(s@left),max(s@left),len=50))curve(dnorm(x,mean=0,sd=2463),add=T,col='blue',lwd=3)
Image Handling
Reading image
install.packages(c('png','jpeg')) # installing packages required for image manipulation
library(png) # importing packages to manipulate images
library(jpeg)
x = readPNG('image_path.png') # reading image data and storing in an object (variable)
y = readJPEG('gorkhey.jpg')
dim(x) # 648 1042 3
str(x) # num [1:648, 1:1042, 1:3] 0.71 0.71 0.71 0.71 0.71 …
648 is no of rows and 1042 is no of columns while 1,2,3 signifies red,green,blue component of a specific pixel. There is (can be) a 4th component alpha that signifies the transparency level of the pixel from 0(invisible) to 1(completely visible)
Plotting image
plot(as.raster(x))
Functions to create a raster object (representing a bitmap image) and coerce other objects to a raster object. An object of class "raster" is a matrix of colour values as given by rgb representing a bitmap image.
example:-
plot(as.raster(x),xlim=c(105,110),ylim=c(80,86),inter=F) # plotting specific selected part of image, inter = interpolation , default TRUE gives smooth out(blur) image
x[1,1,] # gives all structure of 1st row 1st column pixel ; 0 0 0 1 (black)
plot(as.raster(x[,,1])) #should gives all row all column red colour , but don’t as 1==red but in presence of no other we can’t differentiate between colors so we get black&white image, to avoid this
Image manipulation
x[,,2] = 0 #green component 0
x[,,3] = 0 #blue component 0
x[,,4] = 1 #transparency 1
plot(as.raster(x)) #now you will get the required red parts of image only
x[,,1] = x[,,1]/2 #reducing the red component to half
plot(as.raster(x))
p = locator(1)
| |
x[p$y,p$x,] # should give structure of the selected p, but don’t as we wrote p$y,p$x, and not p$x,p$y, as we choose row then column but for locator x axis is horizontal one so it give the column and same argument for y axis so it starts from bottom to top of image while we need opposite and hence we got the data of wrong point (mirror image in mid row)
x[144 - p$y,p$x,] # so to correct that we subtract p$y from no of rows (=144) to get data of selected point
Creating an image
example:-
y = array(0,dim=c(100,200,3)) # initailly 0, three dimensional (100 * 200 * 3) array
y[,,1:2] = 1 # red = green = 1
plot(as.raster(y)) # you will get 100*200 yellow(r+g) image
Try yourself
TRY TO UNDERSTAND THE FOLLOWING R CODE (with reference to what you have learned above) and run them.
| |
| |
| |
| |
| |
| |
| |
| |
| |
Please report any mistake/error or doubt via mail.